ABOUT BEAt-DKD
Diabetic kidney disease (DKD) is the leading cause of end stage renal disease (ESRD), and its global incidence and prevalence have reached epidemic dimensions in recent years. Unfortunately, there are no effective means to prevent or cure DKD. There are many potential reasons for these shortcomings:
- The current diagnosis of diabetes based on the measurement of one metabolite, glucose, is inaccurate and not very useful in predicting disease outcome and choice of therapy.
- Few, if any, studies have taken into account the consequences of early dysglycemia for development of DKD (also referred to as metabolic memory).
- The diagnosis of DKD is usually not based on histological analysis, which is the only way to differentiate DKD from chronic kidney disease (CKD) of other aetiology in patients with diabetes.
- Not much progress has been made in the identification of novel biomarkers for DKD, and micro-/macroalbuminuria still remains among the best predictors of DKD in addition to the decline in glomerular filtration rate (GFR or eGFR).
Only few studies have attempted to explore the full potential of urine and plasma as sources of kidney biomarkers and even fewer have been able to associate such markers with kidney biopsy analyses. Development of novel therapies will require a better understanding of the pathways leading to DKD and better biomarkers to monitor disease progression and treatment responses.
The overall goals of the Biomarker Enterprise to Attack DKD (BEAt-DKD) consortium are:
to provide a holistic systems medicine view of the pathogenesis of DKD with the aim to identify targetable mechanisms and pathways underlying initiation and progression of DKD, applying a novel sub-classification of diabetes,
to identify and validate biomarkers of disease progression and treatment responses representing first steps towards precision medicine in the management of DKD.
SCIENTIFIC WORK PACKAGES
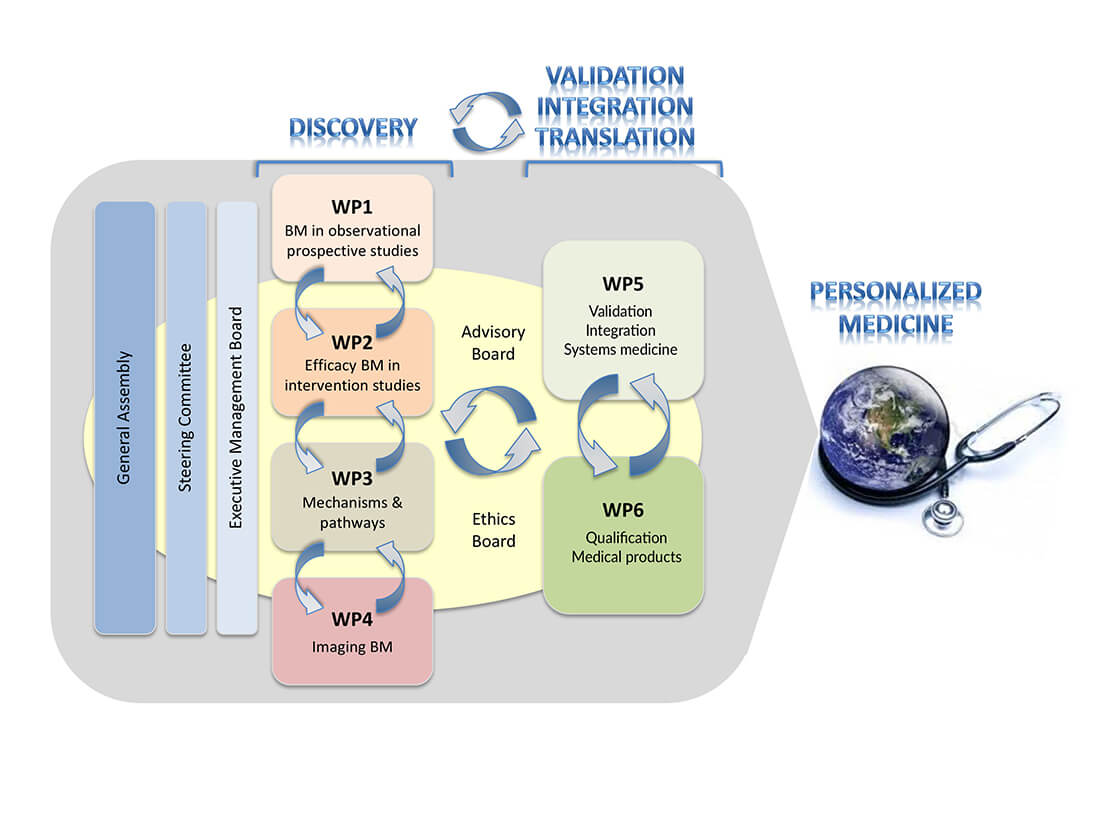
BEAt-DKD is organized into four “discovery” Work Packages (WP1-4) that feed their findings into two “validation, integration and translation” work packages (WPs 5-6). All efforts focus on the discovery, validation and qualification of prognostic, predictive and safety DKD biomarkers to allow improved patient stratification for use in innovative clinical trials aiming at prevention and better management of established DKD, i.e. steps towards precision medicine: “The right prevention and treatment, to the right patients at the right time”.
Biomarkers in observational prospective studies (WP1)
KEY OBJECTIVES OF WP1
- To identify and validate new serum/plasma and urinary biomarkers from, and applicable to, broad populations in order to differentiate fast from slow progressors of DKD in classical T1D and T2D, as well as to explore whether these biomarkers are enriched in some of the new clusters obtained by using a more fine-tuned diabetes classification.
- To extend existing prospective cohorts of DKD patients to follow and collect periodic longitudinal data over several years, focusing on CKD stages 2-4.
WP1 on one hand will use cutting edge technology to discover novel biomarkers, including exploration of the wealth of information carried by urinary vesicles and urine sediment, but will also use established tools for the validation of biomarker candidates which were recently identified by BEAt-DKD partners (i.e. those from IMI SUMMIT, SYSKID) and which have shown excellent performance in pilot studies. This approach will guarantee us a head start with the validation of existing markers in study cohorts specifically designed for the study of DKD with a wide spread of CKD stages and at the same time, allow partners to work on the discovery of novel markers during the first 3 years.
Lund University, University of Helsinki, University of Eastern Finland, University Medical Center Freiburg, University Clinic Erlangen, Medizinische Universität Innsbruck, University Hospital Regensburg, Medical University of Vienna, Lipotype, JDRF, Sanofi-Aventis, Astellas, Eli Lilly, Bayer Pharma, Boehringer Ingelheim, University Medical Center Hamburg-Eppendorf
Efficacy biomarkers in intervention studies (WP2)
KEY OBJECTIVES OF WP2
- Identify and validate a set of biomarkers that can be used to predict drug response before the patient is exposed to treatment (predictive biomarkers) to allow for optimal stratified design of clinical trials, as well as biomarkers which change with standard of care or novel therapies in order to monitor drug efficacy (dynamic biomarkers)
- Integrate the short-term (i.e. weeks to months) effects of a drug on multiple biomarkers in order to optimally predict a drug effect on renal (and cardiovascular) outcomes.
- Integrate the predictive and dynamic biomarkers with clinical phenotype information and drug responses in biochemical and physical parameters in order to (1) obtain a better understanding of the underlying molecular mechanisms that determine individual differences in therapy response; (2) provide useful insights into potential new drug targets.
WP2 focuses on the discovery of biomarkers that predict if a patient with DKD responds to a given drug. In order to identify predictive biomarkers and dynamic biomarkers, the pharmacokinetic/genetic background should be considered. In addition, drug response variability can also be attributed to variability in underlying individual molecular mechanisms driving DKD progression. In this respect, biomarkers should be identified to indicate whether (1) the patient is at risk of progression and (2) which biological process(es) are activated in a specific patient and then (3) match the appropriate treatment as reflected by the drug´s molecular mode of action to the individual’s underlying molecular pathophysiology. WP2 will not only focus on single predictive/dynamic biomarkers but will combine them into drug response classifiers. Predictive biomarkers will be established in cooperation with WP5 and will serve as tools for baseline enrichment strategies.
The WP2 team will conduct experimental studies in cells and animals as well as use blood and urine samples from patients from already finished trials, to determine which molecular pathways are targeted by the drugs of interest. Advanced systems biology methods will generate molecular mechanism of action models, which are complemented with actual kidney tissue data from rat and mice kidney tissues. The generated data in all these studies will be combined to maximize the chance of discovering the best biomarker for drug response prediction. The discovered biomarkers will then be measured in new clinical studies and in clinical trials to verify that their performance in predicting drug response is truly correct.
Lund University, University of Helsinki, University Medical Center Groningen, University of Dundee, Istituto di Ricerche Farmacologiche Mario Negri, University Clinic Erlangen, Medizinische Universität Innsbruck, Medical University of Vienna, Lipotype, Sanofi-Aventis, Astellas, Eli Lilly, Bayer Pharma, Boehringer Ingelheim
Mechanisms and pathways (WP3)
KEY OBJECTIVES OF WP3
- Identify targetable mechanisms and pathways underlying initiation and progression of DKD in order to predict human biomarkers that can subsequently be tested in the clinical materials by the other WPs
- Back-translate clinical results from WP1, 2, 4 and 5 to customized in-vitro and in-vivo models for development of rapid validation tests and HighThroughPut (HTP) techniques.
WP3 will capitalize on cutting edge technology and beyond state-of-the-art model systems provided by its partners. WP3 partners have vast expertise in the development and use of novel genetic, epigenetic, biochemical and physiological experimental tools and approaches, including validated animal models of DKD, to select or set up preclinical model systems for translational studies. It will validate key discoveries from WP1 and WP2 in a variety of in vitro human and in vitro and in vivo animal models of DN and elucidate potential mechanisms by which the “biomarker molecules” may be driving nephropathy progression. The team will also interrogate key molecular pathways involved in the progression of renal disease in animal and human diabetic models, which will feed back into WP1 for interrogation in the human cohorts to determine if these are also important here. Together this unique BEAt-DKD experimental platform will combine highly efficient renal cell isolation protocols, molecular fingerprinting, genome editing, cell culture systems, transgenic mice models and model organisms to systematically validate, identify and predict biomarkers and novel druggable pathways in DKD. Results from WP3 efforts using cell/animal experimental models will be integrated in WP5 and in this way, systematically connected to the human studies.
Lund University, University Medical Center Freiburg, University of Bristol, University of Hull, University of Bari, AbbVie, Sanofi-Aventis, Astellas, Eli Lilly, Bayer Pharma, Novo Nordisk, Boehringer Ingelheim, University Medical Center Hamburg-Eppendorf
Development, identification and validation of prognostic and predictive imaging biomarkers (WP4)
KEY OBJECTIVES OF WP4
- Identify imaging biomarkers that improve the prognosis in DKD, in particular the prediction of functional decline and the differentiation of fast from slow progressors. This will address the need for improved patient segmentation in clinical trials, aiming to reduce cost and duration of clinical trials by focusing on the patient segment where treatment effects are most likely to become apparent in a limited time frame.
- Identify imaging biomarkers of disease progression that have the potential to detect treatment-induced changes in the disease trajectory before changes in glomerular filtration rate become apparent. This will address the need for better end-points for small, rapid and informed trials staged early in the drug development lifecycle aiming to reduce the risk of expensive late-stage trial failures.
The central task in WP4 is a longitudinal 4-year observational study in 500 patients with early stage DKD, recruited in 5 sites across Europe (Leeds, Bari, Turku, Exeter, Bordeaux) with central biobanking in Lund. All patients will be extensively characterised at baseline with clinical assessments, demographics and medical histories, blood/plasma and urine samples, and standardised MRI & US protocols. A cross-sectional analysis of these baseline data will aim to identify associations between imaging biomarkers and known biomarkers of disease progression. As a secondary objective, patients will be followed up annually to determine whether imaging biomarkers collected at baseline improve predictions of functional decline. WP4 will also perform ancillary sub-studies in each of the 5 sites to gain additional insights in the role and interpretation of imaging biomarkers: comparisons against gold-standard PET methods (Turku), validation against histopathology (Bari), correlation against microvascular assessments (Exeter), detection of disease progression (Bordeaux) and development of novel imaging biomarkers (Leeds).
Lund University, University of Exeter, University of Turku, University of Leeds, Hopitaux de Bordeaux, University of Bari, University of Michigan
Integration and prioritization of DKD biomarkers and targets
(systems medicine; WP5)
KEY OBJECTIVES OF WP5
- Develop a project specific data management system to support joint analysis of distributed data
- Integrate data on candidate DKD biomarkers and drug targets using systems biology approaches
- Access and integrate additional data that supports the further validation and refinement of candidate biomarkers and drug targets for DKD
- Generate lists of candidate biomarkers that can guide direct assessment in WPs 1-4
- Generate refined lists of “high quality” validated prognostic/predictive DKD biomarkers and drug targets that can be forwarded to WP6 for further assessment
WP5 will integrate data on multiple candidate biomarkers and treatment targets arising from efforts in WPs 1-4, plus those gleaned from existing and external studies, and, taking advantage of the diverse expertise within BEAt-DKD, set these in their functional context.
The first WP5 actions centre on the development and implementation of a Data Management Plan, including set up of a federated database (to manage access to clinical data that cannot be shared across sites) connected to the central BEAt-DKD database at SIB that will collect genomic and other datasets from across BEAt-DKD as well as other publicly available data sets. Individual clinical datasets are being harmonized using CDISC ontologies, which has been implemented across several BEAt-DKD sites. This WP has close collaborations with WP1 to prioritize biomarkers for inclusion in their respective data acquisition efforts, distilling a priority list of potential biomarkers for DKD from the efforts of previous projects such as IMI SUMMIT and SYSKID, and with WP2 to enhance understanding of the molecular consequences of drug administration on the kidney, and support ongoing biomarker prioritization, through acquisition of transcriptomic data from a range of cellular and rodent models exposed to pharmaceuticals of interest.
WP5 will therefore serve as an integration tool ensuring that the final set of actionable biomarkers and targets benefits from the widest input in terms of domain expertise, background data, biosamples and analytical proficiency.
Lund University, University Medical Center Groningen, University of Oxford, University of Dundee, Medizinische Universität Innsbruck, University of Michigan, Swiss Institute of Bioinformatics, JDRF, Sanofi-Aventis, Eli Lilly, Novo Nordisk
Optimization of trial design and preparation of implementation in the regulatory process (WP6)
KEY OBJECTIVES OF WP6
- Use biomarker scores to optimize the design of trials that test the effectiveness of drugs for renal protection in diabetic nephropathy. This will comprise optimal patient selection and randomization, so called enrichment, with respect to renal risk of the patient and the drug response of the patient. To this end, baseline biomarkers that predict renal risk and baseline as well as biomarkers that predict renal response (including target engagement and dose response) will be used as they are developed by WPs 1-4 and integrated and validated by WP5.
- Interact with the regulatory agencies European Medicine Agency (EMA) and Food and Drug Administration (FDA) to discuss these novel approaches and design, to be integrated into the registration and qualification process
The overarching goal of WP6 is to optimize the trial design towards precision medicine and integration in the regulatory process of drug registration, using available and newly developed biomarker sets and ongoing trials. The specific goals of WP6 are to support engagement with regulators toward potential qualification of biomarker/biomarker panels.
BEAt-DKD’s WP6 focuses on improving development of new treatments in DKD by applying a personalized medicine approach to clinical study design. Personalized medicine identifies the right treatment for the right patient with the help of respective markers. WP6 aims to achieve this by developing a blueprint for such clinical studies by incorporating a multiple biomarker score into a study protocol template for testing the efficacy and safety of new treatments in DKD. WP6 holistically integrates the knowledge generated from the other WPs in the BEAt-DKD project. Implementing a new biomarker tool in clinical research requires a close collaboration of numerous key stakeholders. Therefore, WP6 is engaging with patient and physician groups, regulators, health technology assessors, and scientists from academia, pharmaceutical, and biotechnology companies to achieve its goals. WP6 holds focused meetings with these stakeholders to identify opportunities and challenges for implementing new biomarker tools in clinical practice in order to move personalize medicine forward for better prevention, management, and improved outcomes in DKD.
Lund University, University Medical Center Groningen, University of Oxford, Medizinische Universität Innsbruck, University of Michigan, JDRF, AbbVie, Astellas, Bayer Pharma, Novo Nordisk, Boehringer Ingelheim